Use of Linear and Logistic Regression Coefficients with Lasso (L1) and Ridge (L2) Regularization for Feature Selection in Machine Learning
Watch Full Playlist: https://www.youtube.com/playlist?list=PLc2rvfiptPSQYzmDIFuq2PqN2n28ZjxDH
Linear Regression
Let’s first understand what exactly linear regression is, it is a straight forward approach to predict the response y on the basis of different prediction variables such x and ε. . There is a linear relation between x and y.
𝑦𝑖 = 𝛽0 + 𝛽1.𝑋𝑖 + 𝜀𝑖
y = dependent variable
β0 = population of intercept
βi = population of co-efficient
x = independent variable
εi= Random error
Basic Assumptions
- Linear relationship with the target y
- Feature space X should have gaussian distribution
- Features are not correlated with other
- Features are in same scale i.e. have same variance
Lasso (L1) and Ridge (L2) Regularization
Regularization is a technique to discourage the complexity of the model. It does this by penalizing the loss function. This helps to solve the overfitting problem.
- L1 regularization (also called Lasso)
Itshrinksthe co-efficients which are less important tozero. That means withLasso regularizationwe canremovesome features. - L2 regularization (also called Ridge)
It does’treducethe co-efficients tozerobut it reduces theregression co-efficientswith this reduction we can identofy which feature has more important. - L1/L2 regularization (also called Elastic net)
A regression model that uses L1 regularization technique is called Lasso Regression and model which uses L2 is called Ridge Regression.
What is Lasso Regularisation
3 sources of error
- Noise:
We can’t do anything with the noise. Let’s focus on following errors. - Bias error:
It is useful to quantify how much on anaverageare the predicted values different from the actual value. - Variance:
On the other side quantifies how are the prediction made on thesame observationdifferent from each other.
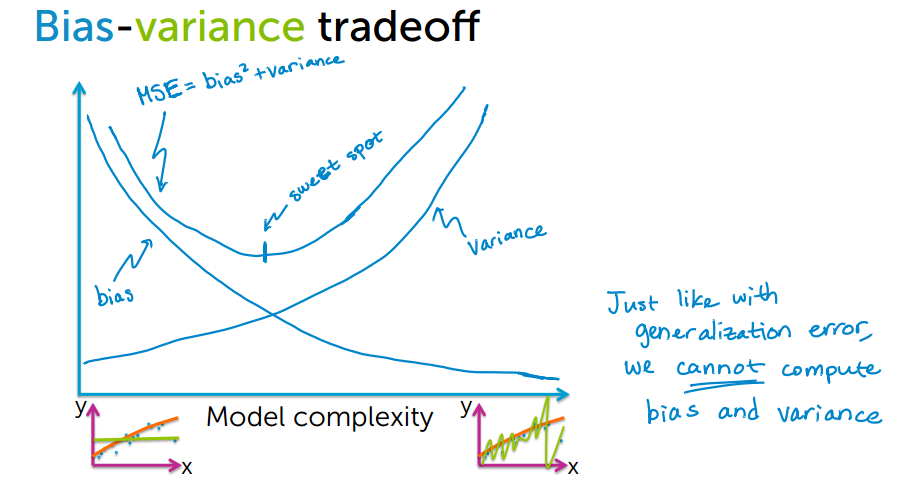
Now we will try to understand bias - variance trade off from the following figure.
By increasing model complexity, total error will decrease till some point and then it will start to increase. W need to select optimum model complexity to get less error.
If you are getting high bias then you have a fair chance to increase model complexity. And otherside it you are getting high variance, you need to decrease model complexity that’s how any machine learning algorithm works.
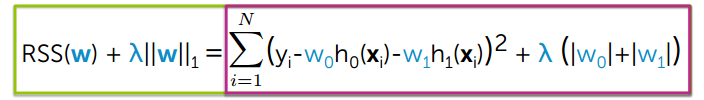
The L1 regularization adds a penalty equal to the sum of the absolute value of the coefficients.
We can observe from the following figure. The L1 regularization will shrink some parameters to zero. Hence some variables will not play any role in the model to get final output, L1 regression can be seen as a way to select features in a model.
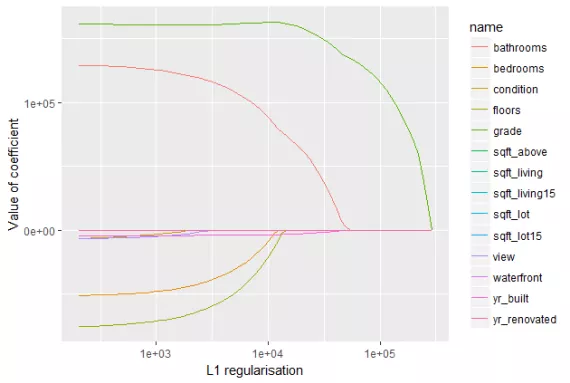
Let’s observe the evolution of test error by changing the value of λ from the following figure.
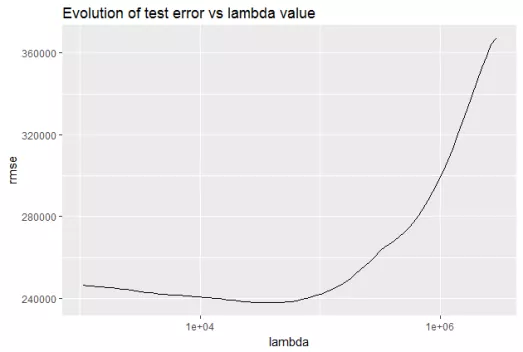
How to choose λ
Let’s move ahead and choose the best λ.
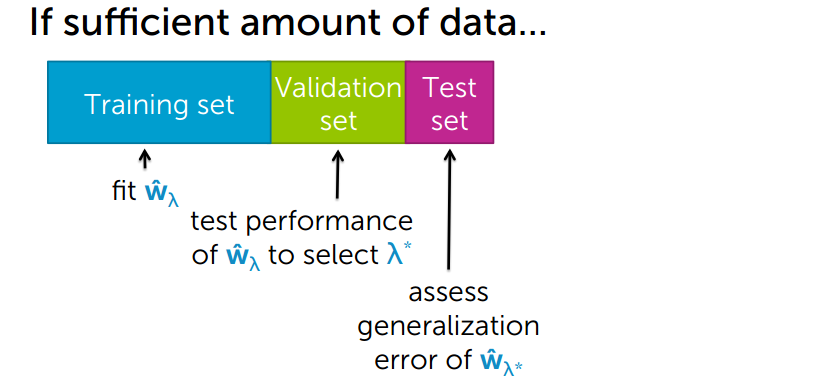
We have a sufficient amount of data. In that we can split our data into 3 sets those are
- Training set
- Validation set
- Test set
- In the training set, we fit our model and set
regression co-efficientswith theregularization. - Then we test our model’s
performanceto select λ
- on
validation set, if any thing wrong with the model likeless accuracywe validate on thevalidation setthen we change the parameter the we go back to thetraining setand so the optimization. - Finally, it will do generalize testing on the
test set.
What is Ridge Regularisation
Let’s first understand what exactly Ridge regularization:
The L2 regularization adds a penalty equal to the sum of the squared value of the coefficients.
λ is the tuning parameter or optimization parameter.
w is the regression co-efficient.
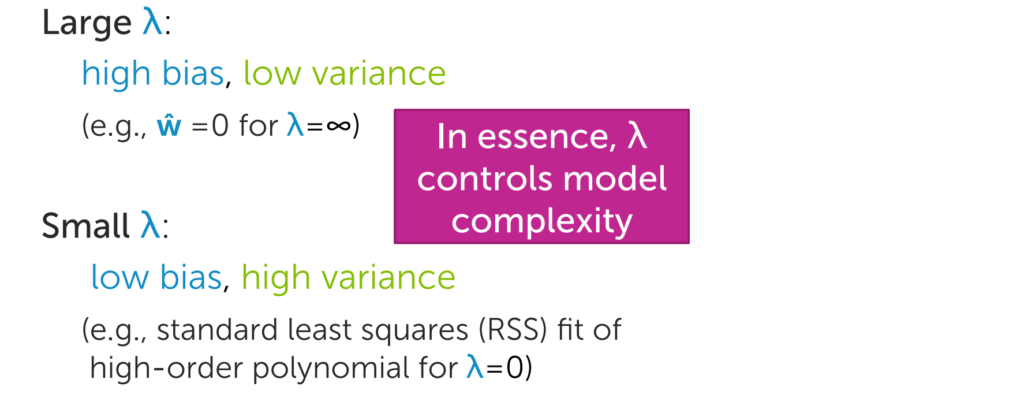
In this regularization,
if λ is high then we will get high bias and low variance.
if λ is low then we will get low bias and high variance
So what we do we will find out the optimized value of λ by tuning the parameters. And we can say λ is the strength of the regularization.

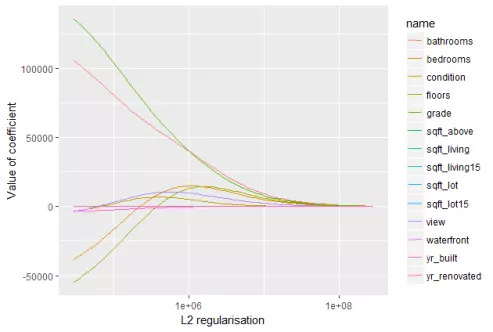
The L2 regularization will force the parameters to be relatively small, the bigger the penalization, the smaller (and the more robust) the coefficients are.
Difference between L1 and L2 regularization
Let’s discuss the difference between L1 and L2 regularization:
| L1 Regularization | L2 Regularization |
|---|---|
| It penalizes sum of absolute value of weights | It regularization penalizes sum of square weights |
| It has a sparse solution | It has a non sparse solution |
| It has multiple solutions | It has one solution |
| It has built in feature selection | It has no feature selection |
| It is robust to outliers | It is not robust to outliers |
| It generates model that are simple and interpretable but cannot learn complex patterns | It gives better prediction when output variable is a function of all input features |
Load the dataset
Loading required libraries
import numpy as np import pandas as pd import seaborn as sns import matplotlib.pyplot as plt %matplotlib inline
from sklearn.model_selection import train_test_split from sklearn.ensemble import RandomForestClassifier from sklearn.feature_selection import SelectKBest, SelectPercentile from sklearn.metrics import accuracy_score
from sklearn.linear_model import LinearRegression, LogisticRegression from sklearn.feature_selection import SelectFromModel
titanic = sns.load_dataset('titanic')
titanic.head()| survived | pclass | sex | age | sibsp | parch | fare | embarked | class | who | adult_male | deck | embark_town | alive | alone | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 3 | male | 22.0 | 1 | 0 | 7.2500 | S | Third | man | True | NaN | Southampton | no | False |
| 1 | 1 | 1 | female | 38.0 | 1 | 0 | 71.2833 | C | First | woman | False | C | Cherbourg | yes | False |
| 2 | 1 | 3 | female | 26.0 | 0 | 0 | 7.9250 | S | Third | woman | False | NaN | Southampton | yes | True |
| 3 | 1 | 1 | female | 35.0 | 1 | 0 | 53.1000 | S | First | woman | False | C | Southampton | yes | False |
| 4 | 0 | 3 | male | 35.0 | 0 | 0 | 8.0500 | S | Third | man | True | NaN | Southampton | no | True |
titanic.isnull().sum()
survived 0 pclass 0 sex 0 age 177 sibsp 0 parch 0 fare 0 embarked 2 class 0 who 0 adult_male 0 deck 688 embark_town 2 alive 0 alone 0 dtype: int64
Let’s remove age and deck from the feature list
titanic.drop(labels = ['age', 'deck'], axis = 1, inplace = True) titanic = titanic.dropna() titanic.isnull().sum()
survived 0 pclass 0 sex 0 sibsp 0 parch 0 fare 0 embarked 0 class 0 who 0 adult_male 0 embark_town 0 alive 0 alone 0 dtype: int64
Let’s get the features of the data
data = titanic[['pclass', 'sex', 'sibsp', 'parch', 'embarked', 'who', 'alone']].copy() data.head()
| pclass | sex | sibsp | parch | embarked | who | alone | |
|---|---|---|---|---|---|---|---|
| 0 | 3 | male | 1 | 0 | S | man | False |
| 1 | 1 | female | 1 | 0 | C | woman | False |
| 2 | 3 | female | 0 | 0 | S | woman | True |
| 3 | 1 | female | 1 | 0 | S | woman | False |
| 4 | 3 | male | 0 | 0 | S | man | True |
data.isnull().sum()
pclass 0 sex 0 sibsp 0 parch 0 embarked 0 who 0 alone 0 dtype: int64
sex = {'male': 0, 'female': 1}
data['sex'] = data['sex'].map(sex)
data.head()| pclass | sex | sibsp | parch | embarked | who | alone | |
|---|---|---|---|---|---|---|---|
| 0 | 3 | 0 | 1 | 0 | S | man | False |
| 1 | 1 | 1 | 1 | 0 | C | woman | False |
| 2 | 3 | 1 | 0 | 0 | S | woman | True |
| 3 | 1 | 1 | 1 | 0 | S | woman | False |
| 4 | 3 | 0 | 0 | 0 | S | man | True |
ports = {'S': 0, 'C': 1, 'Q': 2}
data['embarked'] = data['embarked'].map(ports)
who = {'man': 0, 'woman': 1, 'child': 2}
data['who'] = data['who'].map(who)
alone = {True: 1, False: 0}
data['alone'] = data['alone'].map(alone)
data.head()| pclass | sex | sibsp | parch | embarked | who | alone | |
|---|---|---|---|---|---|---|---|
| 0 | 3 | 0 | 1 | 0 | 0 | 0 | 0 |
| 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 |
| 2 | 3 | 1 | 0 | 0 | 0 | 1 | 1 |
| 3 | 1 | 1 | 1 | 0 | 0 | 1 | 0 |
| 4 | 3 | 0 | 0 | 0 | 0 | 0 | 1 |
Let’s read the data into x
X = data.copy() y = titanic['survived'] X.shape, y.shape
((889, 7), (889,))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 43)
Estimation of coefficients of Linear Regression
First we will go with estimation of coefficients of linear regression
sel = SelectFromModel(LinearRegression())
Let’s go ahead and fit the model
sel.fit(X_train, y_train)
SelectFromModel(estimator=LinearRegression(copy_X=True, fit_intercept=True, n_jobs=None,
normalize=False),
max_features=None, norm_order=1, prefit=False, threshold=None)With these we have trained our model
Let’s see the features of the model
sel.get_support()
array([ True, True, False, False, False, True, False])
Now we will try to get the co-efficients
sel.estimator_.coef_
array([-0.13750402, 0.26606466, -0.07470416, -0.0668525 , 0.04793674,
0.23857799, -0.12929595])Let’s get the mean of the co-efficients
mean = np.mean(np.abs(sel.estimator_.coef_)) mean
0.13727657291370804
And calculate the absolute value of the co-efficients
np.abs(sel.estimator_.coef_)
array([0.13750402, 0.26606466, 0.07470416, 0.0668525 , 0.04793674,
0.23857799, 0.12929595])features = X_train.columns[sel.get_support()] features
Index(['pclass', 'sex', 'who'], dtype='object')
Let’s get the transformed version of x_train and x_test.
X_train_reg = sel.transform(X_train) X_test_reg = sel.transform(X_test) X_test_reg.shape
(267, 3)
Let’s implement the randomForest function and we wil do the training of the model.
def run_randomForest(X_train, X_test, y_train, y_test):
clf = RandomForestClassifier(n_estimators=100, random_state=0, n_jobs=-1)
clf = clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))%%time run_randomForest(X_train_reg, X_test_reg, y_train, y_test)
Accuracy: 0.8239700374531835 Wall time: 250 ms
Now we will get the accuracy and wall time on the original data set
%%time run_randomForest(X_train, X_test, y_train, y_test)
Accuracy: 0.8239700374531835 Wall time: 252 ms
X_train.shape
(622, 7)
Logistic Regression Coefficient with L1 Regularization
Let’s move ahead with the L1 regularization to select the features.
sel = SelectFromModel(LogisticRegression(penalty = 'l1', C = 0.05, solver = 'liblinear')) sel.fit(X_train, y_train) sel.get_support()
array([ True, True, True, False, False, True, False])
Let’s get the regression co-efficients.
sel.estimator_.coef_
array([[-0.54045394, 0.78039608, -0.14081954, 0., 0. ,0.94106713, 0.]])
Let’s get the transformed version of x_train and x_test by using the function transform()
X_train_l1 = sel.transform(X_train) X_test_l1 = sel.transform(X_test)
Now we will get the accuracy
%%time run_randomForest(X_train_l1, X_test_l1, y_train, y_test)
Accuracy: 0.8277153558052435 Wall time: 251 ms
L2 Regularization
Let’s move ahead with the L2 regularization to select the features.
sel = SelectFromModel(LogisticRegression(penalty = 'l2', C = 0.05, solver = 'liblinear')) sel.fit(X_train, y_train) sel.get_support()
array([ True, True, False, False, False, True, False])
sel.estimator_.coef_
array([[-0.55749685, 0.85692344, -0.30436065, -0.11841967, 0.2435823 ,
1.00124155, -0.29875898]])X_train_l1 = sel.transform(X_train) X_test_l1 = sel.transform(X_test)
Let’s check the accuracy of the data
%%time run_randomForest(X_train_l1, X_test_l1, y_train, y_test)
Accuracy: 0.8239700374531835 Wall time: 250 ms
1 Comment